Pharmacotherapy, which remains the cornerstone of chronic pain management, includes analgesics such as nonsteroidal anti-inflammatory drugs (NSAIDs), the more potent opioids, and adjunct therapies such as antidepressants and anticonvulsants.1 It is well-recognized that interindividual differences exist in the general population in terms of pain perception, response to analgesic treatment, and toxicities experienced.2 This variability may be influenced by age, health status, sex, ethnicity, diet, and other factors. However, it is becoming increasingly evident that a considerable percentage of the variability may be genetically controlled.2 Several such genetic alterations have been characterized in candidate genes encoding molecules important for pain transmission, which result in impaired function of these gene products to ultimately impact the efficacy and side effect profiles of several analgesics, thus defining the active field of pharmacogenomics in pain medicine. The understanding gained from pharmacogenomics holds the potential for the practice of personalized medicine, which will empower healthcare professionals to make rational clinical decisions regarding drug selection, the prediction of therapeutic responses, and the identification of patients at high risk for adverse events. In this article, an overview of prominent genetic alterations underlying interpatient diversity and their clinical relevance in pain management is presented.
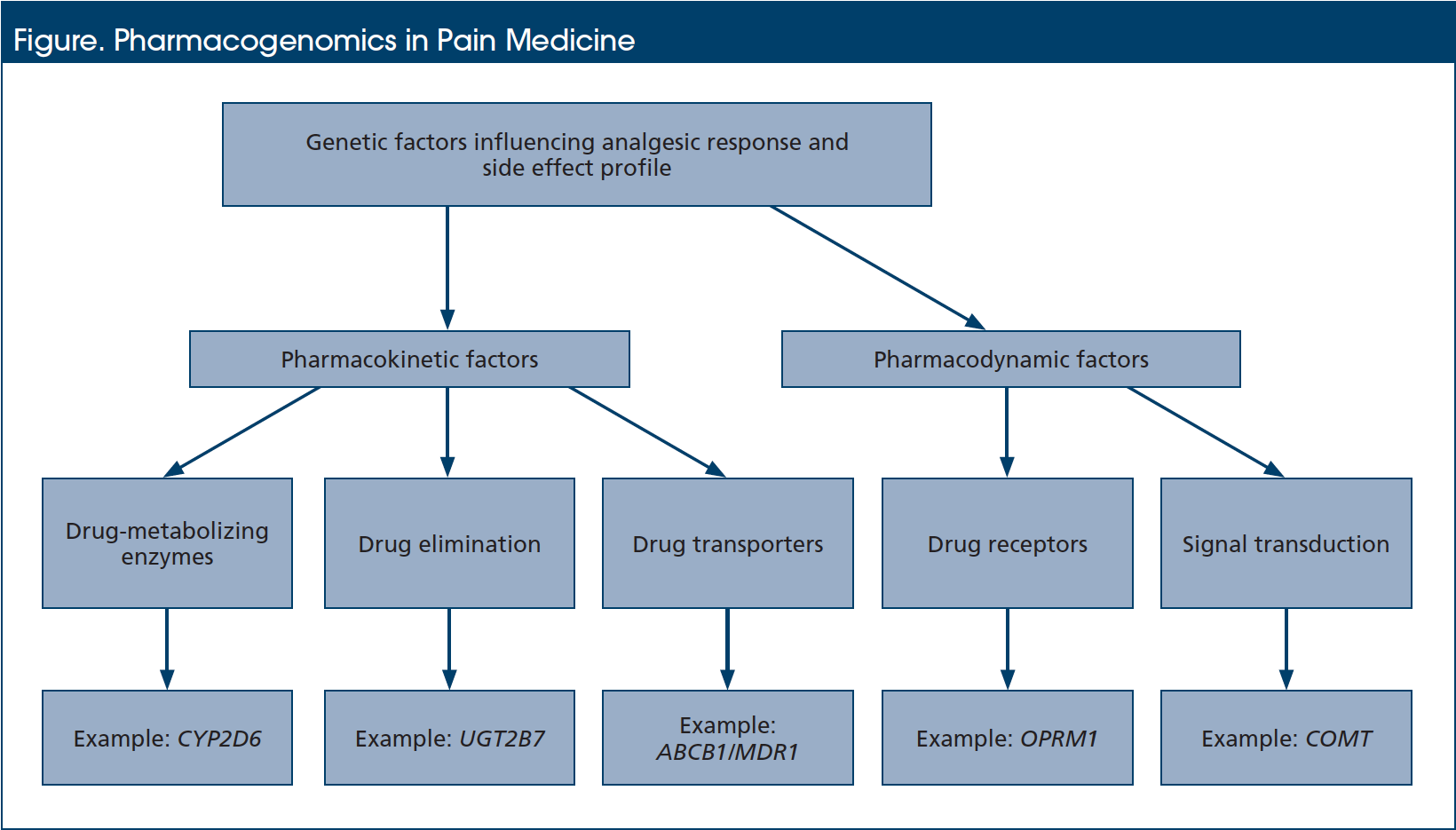
The pharmacogenetic information contributing to interpatient variability in drug response and toxicity may be broadly classified into factors affecting drug pharmacokinetics and factors affecting pharmacodynamics (Figure).3 With regard to analgesics, alterations in pharmacokinetic factors greatly affect drug metabolism, elimination, and/or transport, to influence the relationship between drug dose and steady-state serum drug concentration.3 This is considered to be the primary mechanism contributing to interindividual variability in drug response, and it typically involves genes encoding the drug-metabolizing cytochrome P450 (CYP) family of enzymes, enzymes involved in the glucuronidation metabolic pathway, and drug transporter proteins.4 On the other hand, alterations in pharmacodynamic factors affect drug target receptors and subsequent signal transduction pathways.3
Pharmacokinetic Factors in Pain Medicine
Cytochrome P450 Enzymes
The most important class of drug-metabolizing enzymes for analgesics is CYP, which is mainly responsible for type I reactions, the modification reactions that inactivate or activate an analgesic administered as an inactive or less active prodrug.5 The CYP enzymes comprise a group of proteins that play an important role in the metabolism of several drugs, including those relevant to pain therapy. Genetic alterations have been described in several CYP enzymes; predominant among these are those involving CYP2D6 and CYP3A4.5
CYP2D6 is a well-characterized P450 enzyme that is involved in the metabolism of approximately 25% of all drugs currently in therapeutic use, and it is implicated in the metabolism of several analgesics, including opioids. There are more than 80 unique alleles that have been described for the CYP2D6 enzyme,5 and the different allele combinations yield phenotypic diversity in the general population. Four phenotypes have been identified that are each associated with differential analgesic responses in humans5,6:
- Extensive metabolizers (EMs): individuals who carry 2 wild-type alleles and exhibit normal enzyme activity
- Poor metabolizers (PMs): individuals who carry 2 nonfunctional alleles and have decreased or absent enzyme activity
- Intermediate metabolizers: individuals who have at least 1 reduced functional allele and decreased enzyme activity
- Ultrarapid metabolizers (UMs): individuals who have multiple copies of a functional allele and/or an allele with a promoter mutation and show increased enzyme activity.
In addition to interindividual differences, interethnic differences that are attributable to genetic variations have also been documented.7
The PM phenotype has been characterized in approximately 7% to 10% of the population; these individuals show an increased frequency of CYP2D6*3, CYP2D6*4, CYP2D6*5, or CYP2D6*6 genetic alterations.2,8 The lower enzyme activity renders the PM subgroup at increased risk for drug overdose when the parent compound is responsible for the analgesic effect, or therapeutic failure if the parent compound is a prodrug that needs to be converted to an active metabolite.5 In North America, the UM phenotype occurs at a lower frequency in the population than the PM phenotype5; the UM genotype may be contributed by duplications of CYP2D6*1, CYP2D6*2, CYP2D6*35, or CYP2D6*41 alleles.6 Higher expression of any of these functional CYP2D6 alleles results in excessive production of the active metabolites, which leads to either therapeutic failure if the parent compound is the active entity or increased exposure and more severe adverse effects if the metabolite produces the analgesic effect.5
Clinical evidence has shown that the plasma concentration of a popular opioid and its metabolite is low in PMs, which directly correlated with the suboptimal analgesic effect of the drug in these individuals.2,6,9 In contrast, the analgesic effect of the same opioid was more pronounced with a higher side effect profile in UMs, because of the higher drug plasma concentration.2,6 This has been corroborated in reports of fatalities among patients with UM phenotypes, particularly infants.10,11 However, it must be mentioned that certain other analgesics were found not to be modulated by CYP2D6 activity; therefore, CYP2D6 genetic alterations may not be of clinical relevance to the analgesic effects of such drugs.12
Genetic variants in the CYP2C family of CYP enzymes, comprising CYP2C8, CYP2C9, CYP2C18, and CYP2C19, have also been shown to have clinical importance in pain management, since they are implicated in the metabolic activation of NSAIDs, among other therapeutically useful drugs.13 Approximately, 42 alleles of CYP2C9 have been identified, with CYP2C9*2 and CYP2C9*3 being the most common variants.13 Such genetic variants have been associated with decreased enzyme activity toward several NSAIDs and selective cyclooxygenase-2 inhibitors.13 In addition, CYP2C9 polymorphic status is the major determinant of systemic drug clearance for several NSAIDs.14
CYP2C8 is another major enzyme that is important for the metabolism of drugs, including NSAIDs.2 A number of clinically relevant polymorphisms of CYP2C8 have been identified, including CYP2C8*2 and CYP2C8*3.2 Moreover, there is strong evidence that the CYP2C8*3 variant is often linked with CYP2C9*2. In a large study of 1468 patients, approximately 96% of those with the CYP2C8*3 allele also carried CYP2C9*2.15 In the case of an NSAID, the combined presence of allelic variants CYP2C9*3, CYP2C9*2, and CYP2C8*3 was found to be a strong determinant of impaired metabolism.16 Current evidence also indicates that individuals harboring the gene variants CYP2C8*3, CYP2C9*2, or CYP2C9*3 are at increased risk for developing acute gastrointestinal bleeding with the use of NSAIDs that are substrates of CYP2C8/9 enzymes.17,18
CYP2C19, another important enzyme in the CYP2C family, is involved in the metabolism of several drugs.13 Approximately 34 allele variants of CYP2C19 have been identified, with CYP2C19*2 and CYP2C19*3 alleles accounting for the majority of the abnormal phenotypes.13 The CYP2C19*2 and CYP2C19*3 variants have been shown to render the enzyme nonfunctional, resulting in decreased drug tolerance and requiring dose adjustments in PMs. In contrast, there is some evidence that the CYP2C19*17 allele supports the UM or EM phenotype.13
CYP3A4 is known to play an important role in the metabolism of some opioids. However, evidence supporting the clinical significance of CYP3A4 genetic variations in modulation of opioid activity has been inconclusive.2 Regardless, some opioid-related fatalities have been attributed to genetic variants of CYP3A4 that contributed to the PM phenotype, leading to impaired drug metabolism.19
Uridine Diphosphate Glucuronosyltransferases
Glucuronidation is a critical metabolic process that leads to the formation of hydrophilic glucuronides and is mediated by a family of enzymes called uridine diphosphate glucuronosyltransferases (UGTs).2 UGT2B7 is a key UGT enzyme that is responsible for the glucuronidation of several analgesics.6 The UGT2B7 gene is also polymorphic, although fewer than 20 allelic variants have been identified. The UGT2B7 C802T allele variant is the most studied.6 In a trial that assessed genetic variations of UGT2B7 in patients receiving an opioid that they controlled themselves, the homozygous 802T/T allele variant yielded the lowest levels of the drug and reduced M6G/opioid ratios compared with the heterozygous C/T and homozygous wild-type C/C genotypes.20
P-Glycoprotein–Mediated Drug Transport
P-glycoprotein (Pgp) is a transporter that is expressed in the liver, kidneys, and blood–brain barrier.6 It is encoded by the ABCB1/MDR1 gene (adenosine triphosphate–binding cassette subfamily isoform-1/multidrug resistance protein 1). Many opioids are known to be Pgp substrates, and their analgesic effects are subject to pharmacogenetic modulation.2 It is hypothesized that functional impairment of Pgp-mediated drug transport may result in increased drug bioavailability, reduced renal clearance, or increased brain concentration of Pgp substrates.6 There are 1425 known polymorphisms of the ABCB1/MDR1 gene, and the C3435T variant is well-known to influence pain relief. The C3435T variant predicted treatment response in 145 patients with cancer, with sensitivity close to 100% and specificity greater than 70%, when combined with a certain polymorphism of the opioid receptor mu 1 (OPRM1) gene.21 In patients receiving spinal analgesia with a specific opioid, presence of the C3435T allele was associated with increased suppression of respiratory rate and need for oxygen.22 In patients exposed to experimental pain and a separate opioid analgesia, the variant alleles C3435T and G2677T were found to be strong determinants of less pronounced adverse drug reactions and increased analgesic effect.23
Pharmacodynamic Factors in Pain Medicine
There are several genetic variations that may impact drug receptor binding and drug target expression levels, such as those that involve opioidreceptors, and those that activate signaling transduction pathways (eg, the enzyme catechol-O-methyltransferase [COMT]).Opioid Receptors
The analgesic effects of opioids are primarily mediated by their agonistic interaction with the mu opioid receptor (MOR), which is encoded by the OPRM1 gene.6 OPRM1 has more than 1800 known polymorphisms; the most prominent among these is polymorphism A118G, which occurs with an allelic frequency of approximately 8% to 17% in Caucasians and nearly 50% in Asians.6 OPRM1 A118G results in an asparagine to aspartic acid substitution at position 40 that removes a glycosylation site, thereby causing altered binding of the endogenous agonist beta-endorphin.24 The functional consequence of the genetic change, as gathered from experimental models, includes lower MOR receptor protein expression, decreased signal transduction following agonist engagement, lower rate of G-protein coupling, and increased inhibitory effects on calcium channels.25-27
In contrast to preclinical data, conflicting findings regarding the association of the A118G polymorphism with opioid requirements and magnitude of pain relief have been reported. One study found that the opioid requirement for pain relief was significantly higher in chronic pain patients homozygous for the wild-type allele compared with that for patients carrying the A118G variant allele.28 Another study found that the polymorphic allele significantly reduced the required median dose of an intrathecal opioid in women receiving labor analgesia.29 This result is directly opposed to findings from other studies, in which an opioid requirement for pain relief was higher in patients homozygous for the 118G allele than in heterozygous and homozygous wild-type carriers.30,31 Reyes-Gibby and colleagues reported that carriers of an OPRM1 mutant genotype required a 93% higher opioid dose than carriers of homozygous wild-type genotype.31 In contrast to these studies, the opioid requirement to achieve pupil constriction, a surrogate measure of opioid functional effect, was unaffected by A118G relative to the wild type, although the polymorphism significantly reduced the potency of the opioid metabolite M6G.32 Moreover, a meta-analysis found no consistent association between the A118G polymorphism and opioid dose requirement, and only a weak association for lower incidence of nausea.33
Catechol-O-Methyltransferase
COMT is a key enzyme involved in the inactivation of catecholamine neurotransmitters, including dopamine, norepinephrine, and epinephrine; it also acts as an important modulator of opioid efficacy.6 Many polymorphisms in the COMT gene have been identified, with the G1947A variant, which results in an amino acid change from valine to methionine at codon 158 (Val158Met), being the best characterized.3 The Val158Met alteration is associated with reduced enzymatic activity resulting from decreased thermal stability of the COMT protein.3
Currently, it is unclear whether the Val158Met variant is associated with variable response to opioid dosing. In a pharmacokinetic study of 207 opioid-treated patients with cancer, COMT Val/Val carriers required a 63% higher dose than carriers of the Met/Met genotype, whereas Val/Met genotype carriers required a 23% higher dose.31 The opioid dose required was lowest in carriers of the COMT Met/Met and OPRM1 AA genotypes. However, results from a large genetic association study of 2294 patients with cancer-related pain, which analyzed polymorphisms in genes with putative influence on opioid effects, such as COMT and OPRM1, did not find any significant association with opioid dosing.34
Conclusions
Based on pharmacogenomic data accumulated thus far, investigators have identified a number of genetic variations in candidate genes that have modest correlations with individual variation in analgesic response and toxicity. A pharmacogenomics-based approach to pain management may allow rational drug selection, resulting in improved treatment efficacy and toxicity profiles. However, in reality, available data supporting the use of identified genetic biomarkers in routine clinical practice to predict an individual’s response to pain medication is unconvincing. Therefore, well-designed prospective studies with robust clinical end points are needed to demonstrate the utility of pharmacogenomics in pain management and to make the concept of personalized medicine a reality.
References
- American Society of Anesthesiologists Task Force on Chronic Pain Management; American Society of Regional Anesthesia and Pain Medicine. Practice guidelines for chronic pain management: an updated report by the American Society of Anesthesiologists Task Force on Chronic Pain Management and the American Society of Regional Anesthesia and Pain Medicine. Anesthesiology. 2010;112:810-833.
- Svĕtlik S, Hronová K, Bakhouche H, et al. Pharmacogenetics of chronic pain and its treatment. Mediators Inflamm. 2013;2013:864319.
- Kapur BM, Lala PK, Shaw JL. Pharmacogenetics of chronic pain management. Clin Biochem. 2014;47:1169-1187.
- Smith HS. Opioid metabolism. Mayo Clin Proc. 2009;84:613-624.
- Leppert W. CYP2D6 in the metabolism of opioids for mild to moderate pain. Pharmacology. 2011;87:274-285.
- Janicki PK. Pharmacogenomics of pain management. In: Deer TR, Leong MS, Buvanendran A, et al, eds. Comprehensive Treatment of Chronic Pain by Medical, Interventional, and Integrative Approaches: The American Academy of Pain Medicine Textbook on Patient Management. New York, NY: Springer Publishing Company; 2013:23-33.
- Masimirembwa CM, Hasler JA. Genetic polymorphism of drug metabolising enzymes in African populations: implications for the use of neuroleptics and antidepressants. Brain Res Bull. 1997;44:561-571.
- Buzkova H, Pechandova K, Slanar O, et al. Frequency of single nucleotide polymorphisms of CYP2D6 in the Czech population. Cell Biochem Funct. 2008; 26:76-81.
- Poulsen L, Brosen K, Arendt-Nielsen L, et al. Codeine and morphine in extensive and poor metabolizers of sparteine: pharmacokinetics, analgesic effect and side effects. Eur J Clin Pharmacol. 1996;51:289-295.
- Madadi P, Ross CJ, Hayden MR, et al. Pharmacogenetics of neonatal opioid toxicity following maternal use of codeine during breastfeeding: a case-control study. Clin Pharmacol Ther. 2009;85:31-35.
- Gasche Y, Daali Y, Fathi M, et al. Codeine intoxication associated with ultrarapid CYP2D6 metabolism. N Engl J Med. 2004;351:2827-2831.
- Wilder-Smith CH, Hufschmid E, Thormann W. The visceral and somatic antinociceptive effects of dihydrocodeine and its metabolite, dihydromorphine: a cross-over study with extensive and quinidine-induced poor metabolizers. Br J Clin Pharmacol. 1998;45:575-581.
- Hirota T, Eguchi S, Ieiri I. Impact of genetic polymorphisms in CYP2C9 and CYP2C19 on the pharmacokinetics of clinically used drugs. Drug Metab Pharmacokinet. 2013;28:28-37.
- Rodrigues AD. Impact of CYP2C9 genotype on pharmacokinetics: are all cyclooxygenase inhibitors the same? Drug Metab Dispos. 2005;33:1567-1575.
- Yasar U, Lundgren S, Eliasson E, et al. Linkage between the CYP2C8 and CYP2C9 genetic polymorphisms. Biochem Biophys Res Commun. 2002;299: 25-28.
- Karazniewicz-Lada M, Luczak M, Glowka F. Pharmacokinetic studies of enantiomers of ibuprofen and its chiral metabolites in humans with different variants of genes coding CYP2C8 and CYP2C9 isoenzymes. Xenobiotica. 2009;39:476-485.
- Blanco G, Martinez C, Ladero JM, et al. Interaction of CYP2C8 and CYP2C9 genotypes modifies the risk for nonsteroidal anti-inflammatory drugs-related acute gastrointestinal bleeding. Pharmacogenet Genomics. 2008;18:37-43.
- Agundez JA, Garcia-Martin E, Martinez C. Genetically based impairment in CYP2C8- and CYP2C9-dependent NSAID metabolism as a risk factor for gastrointestinal bleeding: is a combination of pharmacogenomics and metabolomics required to improve personalized medicine? Expert Opin Drug Metab Toxicol. 2009;5:607-620.
- Jin M, Gock SB, Jannetto PJ, et al. Pharmacogenomics as molecular autopsy for forensic toxicology: genotyping cytochrome P450 3A4*1B and 3A5*3 for 25 fentanyl cases. J Anal Toxicol. 2005;29:590-598.
- Sawyer MB, Innocenti F, Das S, et al. A pharmacogenetic study of uridine diphosphate-glucuronosyltransferase 2B7 in patients receiving morphine. Clin Pharmacol Ther. 2003;73:566-574.
- Campa D, Gioia A, Tomei A, et al. Association of ABCB1/MDR1 and OPRM1 gene polymorphisms with morphine pain relief. Clin Pharmacol Ther. 2008;83:559-566.
- Park HJ, Shinn HK, Ryu SH, et al. Genetic polymorphisms in the ABCB1 gene and the effects of fentanyl in Koreans. Clin Pharmacol Ther. 2007;81:539-546.
- Zwisler ST, Enggaard TP, Noehr-Jensen L, et al. The antinociceptive effect and adverse drug reactions of oxycodone in human experimental pain in relation to genetic variations in the OPRM1 and ABCB1 genes. Fundam Clin Pharmacol. 2010;24:517-524.
- Kroslak T, Laforge KS, Gianotti RJ, et al. The single nucleotide polymorphism A118G alters functional properties of the human mu opioid receptor. J Neurochem. 2007;103:77-87.
- Margas W, Zubkoff I, Schuler HG, et al. Modulation of Ca2+ channels by heterologously expressed wild-type and mutant human micro-opioid receptors (hMORs) containing the A118G single-nucleotide polymorphism. J Neurophysiol. 2007;97:1058-1067.
- Zhang Y, Wang D, Johnson AD, et al. Allelic expression imbalance of human mu opioid receptor (OPRM1) caused by variant A118G. J Biol Chem. 2005;280:32618-32624.
- Oertel BG, Kettner M, Scholich K, et al. A common human micro-opioid receptor genetic variant diminishes the receptor signaling efficacy in brain regions processing the sensory information of pain. J Biol Chem. 2009;284:6530-6535.
- Janicki PK, Schuler G, Francis D, et al. A genetic association study of the functional A118G polymorphism of the human mu-opioid receptor gene in patients with acute and chronic pain. Anesth Analg. 2006;103:1011-1017.
- Landau R, Kern C, Columb MO, et al. Genetic variability of the mu-opioid receptor influences intrathecal fentanyl analgesia requirements in laboring women. Pain. 2008;139:5-14.
- Klepstad P, Rakvag TT, Kaasa S, et al. The 118 A > G polymorphism in the human mu-opioid receptor gene may increase morphine requirements in patients with pain caused by malignant disease. Acta Anaesthesiol Scand. 2004;48:1232-1239.
- Reyes-Gibby CC, Shete S, Rakvag T, et al. Exploring joint effects of genes and the clinical efficacy of morphine for cancer pain: OPRM1 and COMT gene. Pain. 2007;130:25-30.
- Lotsch J, Skarke C, Grosch S, et al. The polymorphism A118G of the human mu-opioid receptor gene decreases the pupil constrictory effect of morphine-6-glucuronide but not that of morphine. Pharmacogenetics. 2002;12:3-9.
- Walter C, Lotsch J. Meta-analysis of the relevance of the OPRM1 118A>G genetic variant for pain treatment. Pain. 2009;146:270-275.
- Klepstad P, Fladvad T, Skorpen F, et al. Influence from genetic variability on opioid use for cancer pain: a European genetic association study of 2294 cancer pain patients. Pain. 2011;152:1139-1145.